Every year food-borne bacteria cause thousands infections in humans and animals. Outbreaks of E.Coli and Salmonella commonly occur in meat, vegetables and processed food products. The most recent and the deadliest recorded E.Coli outbreak caused 39 deaths and over 3500 reported cases of poisoning in May and June 2011 in Germany. An effective method for disease detection and prevention is still missing, mostly due to long incubation times and a high operation cost associated with conventional methods. The most widely used method for bacteria detection, the standard plate count, takes from 24 to 48 hours, due to the time needed for bacteria to grow detectable colonies. This time is too long for most applications in the food industry, especially when food products need to be delivered and sold fresh. Faster methods including PCR or labeled detection and fluorescent imaging are too expensive to be widely applied.
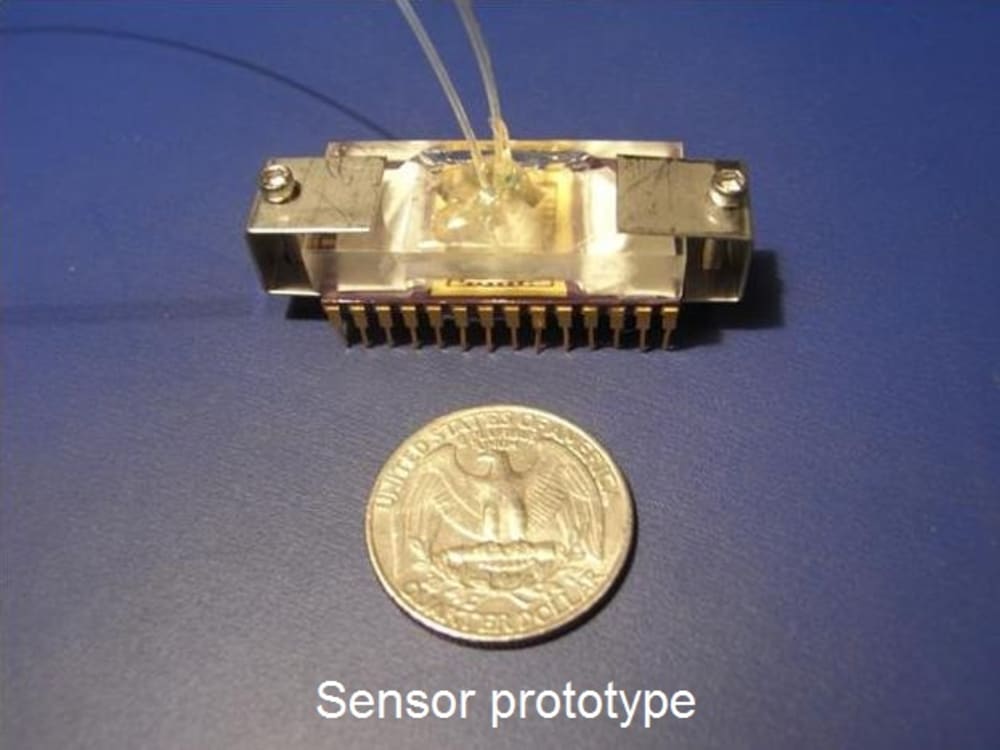
?Screen is the needed MEMS rapid pathogen screener for food-borne illnesses. It has a potential to improve health standards at the consumer level by increasing sensitivity, keeping the cost down and testing time short. This technology will allow screening of up to 100% of food produced in food processing plants before it is delivered to the consumer, thus, for the first time eliminating the risk of infection. ?Screen is portable and ready to use in any conditions. ?Screen is highly accurate, can detect as low as a single bacteria, small, fast, and approximately 50 times cheaper than the established, currently used detection methods. ?Screen can detect any chosen type of bacteria from whole blood or predefined fluid. The device size is less than 1”x1”x0.5”, the cost is as low as 1$ per test.
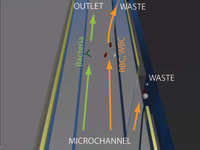
?Screen operation is simple. The user injects a sample of blood into the inlet chamber. Further on, the device processes the sample automatically. Using dielectrophoesis, any present bacteria is separated from the large blood components e.g. red and white blood cells. For bacterial detection we use nanoribbon/nanowire field effect transistors, which are well explored label-free nanosensors, capable of detecting low concentrations of biomolecules i.e. disease markers to provide point-of-care diagnostics [Stern, Vacic et. al Nat. Nanotechnology 2010]. In order to achieve selectivity and specificity of the detection and eliminate false-positive response, the sensors are functionalized with bacteria specific antibodies [Stern, Vacic et. al Small 2010].
?Screen is designed to integrate innovative bacteria separation and demonstrated state of the art detection methods. The whole bacteria separation system has been modeled using the COMSOL Multiphysics software. Sub components for detection have already been successfully prototyped and are currently under testing. The core part of ?Screen is a silicon nanoribbon FET sensor fabricated using CMOS compatible process using optical photolithography and a microfluidic two-step separator for bacteria separation from the whole blood.
?Screen is a miniature device for detecting low number of bacteria under half an hour for only 1$. The ultimate goal is the detection of several hundreds of diseases. It has a potential to revolutionize diagnostics of bacterial diseases.
-
Awards
-
 2011 Grand Prize Winner
2011 Grand Prize Winner
Like this entry?
-
About the Entrant
- Name:Monika Weber
- Type of entry:teamTeam members:Monika Weber, Yale University
Christopher Yerino, Yale University
Hazael Montanaro, Yale University
Kane Siu Lung Lo, Yale University
Mark Reed, Yale University - Software used for this entry:COMSOL Multiphysics, Autodesk Inventor, Blender, L-Edit,
- Patent status:pending