Proteins are large essential macromolecules in the human bodies. Knowing their structure helps the biomedical researchers predict effective drugs. So it is highly important to have a tool for predicting its primary, secondary, tertiary and quaternary structures.
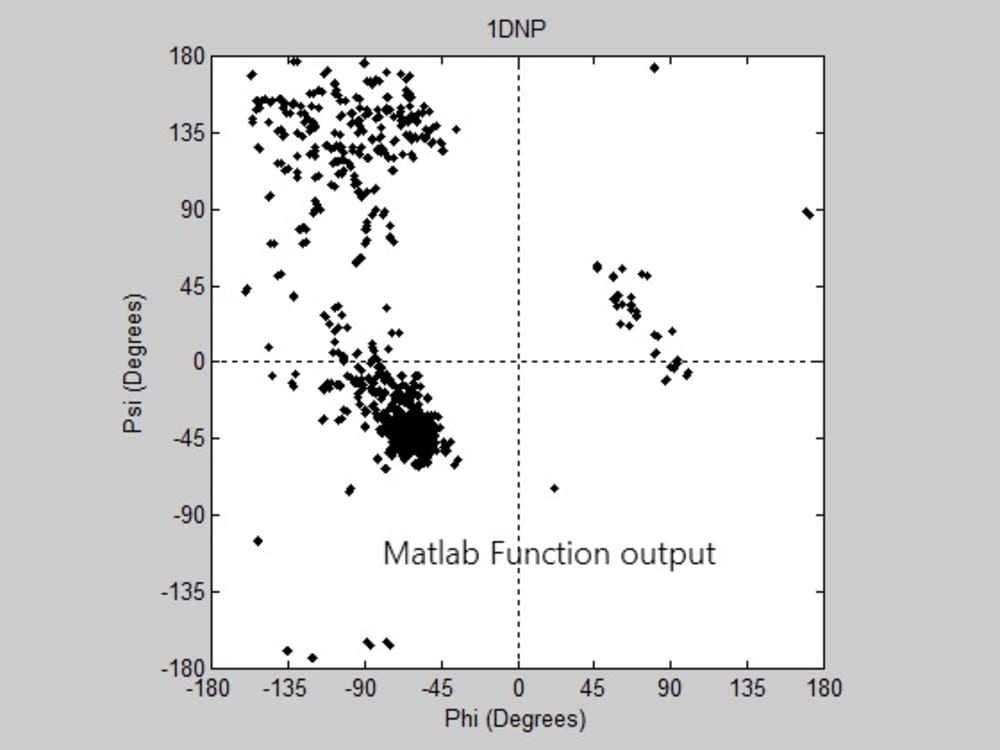
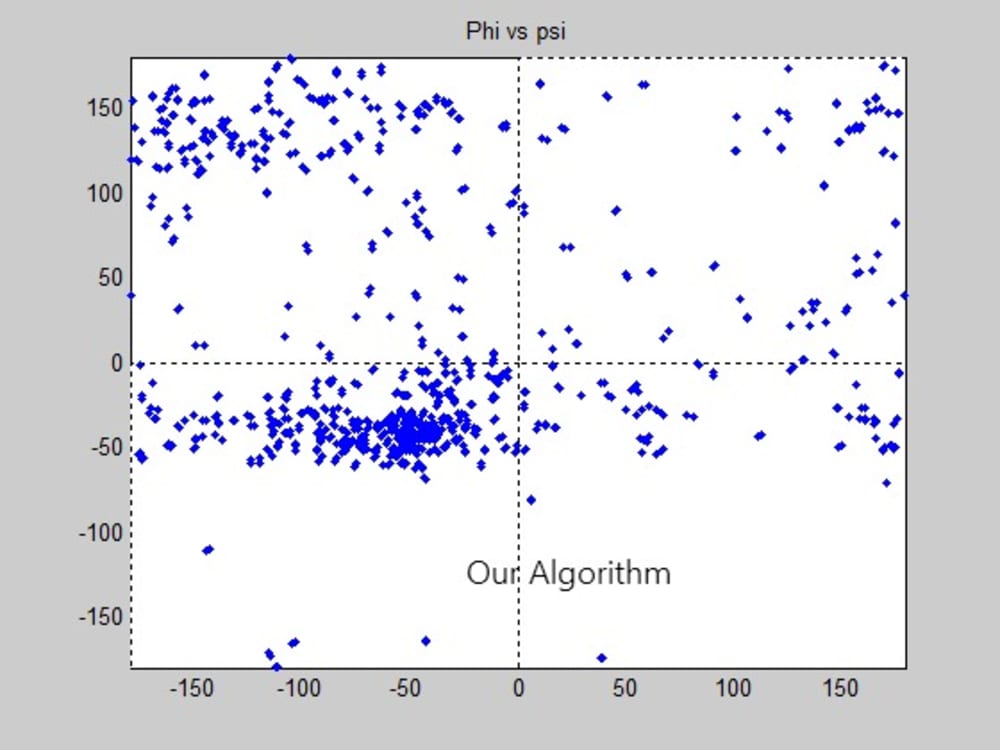
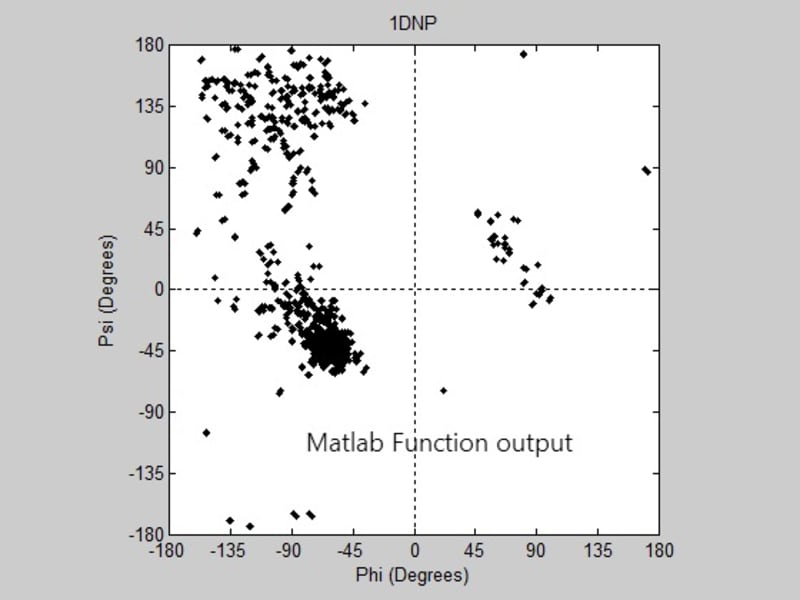
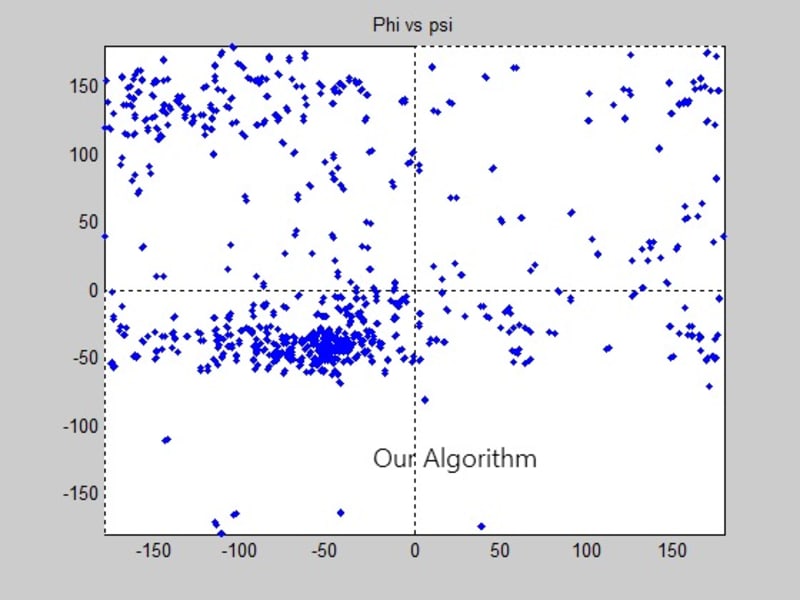
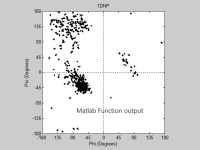
Some protein modeling programs are present nowadays to draw the 3D structure of proteins. However, not all of them show high time and space efficiency. This project introduces a new algorithm for predicting the secondary structure of proteins. It mainly depends on drawing a Ramachandran plot and predicting the sites of the alpha helices and beta sheets, through calculating the angles produced from the rotation of the bonds of the polypeptide backbone, using the normals to the planes on which the atoms of the protein’s backbone lie. The algorithm shows high time efficiency when compared to the built in function for graphing the Ramachandran plot on the MatLab.
Like this entry?
-
About the Entrant
- Name:Essam Radwan
- Type of entry:teamTeam members:Essam Radwan Helmy Hebat-Allah Samy
- Software used for this entry:Matlab
- Patent status:pending